 Armed with an easy-to-use GUI, JASP allows both classical and Bayesian. Residual standard error: 2.033 on 12 degrees of freedom JASP is an open-source statistics program that is free, friendly, and flexible. Lm(formula = Total_amino_acids ~ Genotype * Timepoint, data = AtAAdata) I would like to save these results on my desktop so I don't have to go back and run the tests all over again to see the results! This is what my console looks like: > summary(AtTotal_Model) You need to manually take off the knit_latex() command if you’re knitting to any format other than PDF.I am somewhat new to R, and I don't know how to export the results of a statistical test that I conducted to a text file. column.labels a character vector of labels for columns in regression tables. 
out.header a logical value that indicates whether the LaTeX or HTML le output should contain a code header (if TRUE) or just the chunk of code that creates the output (if FALSE). Importantly, if you knit to HTML now and leave knit_latex() in your code, you won’t see any tables. of the type argument will determine the type of output le. Second, you need to pipe any msummary()s into a special knit_latex() function: msummary(list("A" = model1, "B" = model2, "C" = model3)) %>% knit_latex() Add this header-includes section to your YAML metadata: title: "Whatever" msummary() needs a little help to knit to PDF.įirst, you need to tell the PDF template to use a few LaTeX packages. 
When you use huxreg(), it automatically adapts to whatever output format you’re using (HTML, PDF, Word, etc.). Packageįeed huxreg() a bunch of models: library(huxtable)Īdd column names: huxreg(list("A" = model1, "B" = model2, "C" = model3))ĭownside 2: Some manual work needed to knit to PDF Included below is a helpful summary of the capabilities and limitations of each of these packages, as well as some example code for fixing these limitations. These two functions work slightly differently and you need to make a few minor adjustments depending on how you’re knitting your document. The msummary() function in the modelsummary package.The huxreg() function in the huxtable package.There are two modern R packages that work fairly well for creating these tables: Making these tables with R is generally fairly simple, but you need to do a couple extra steps to make them look nice consistently. If you’re unfamiliar with these kinds of tables, check out this helpful guide to how to read them. It’s often helpful to put the results from regression models in a side-by-side table so you can compare coefficients across different model specifications. 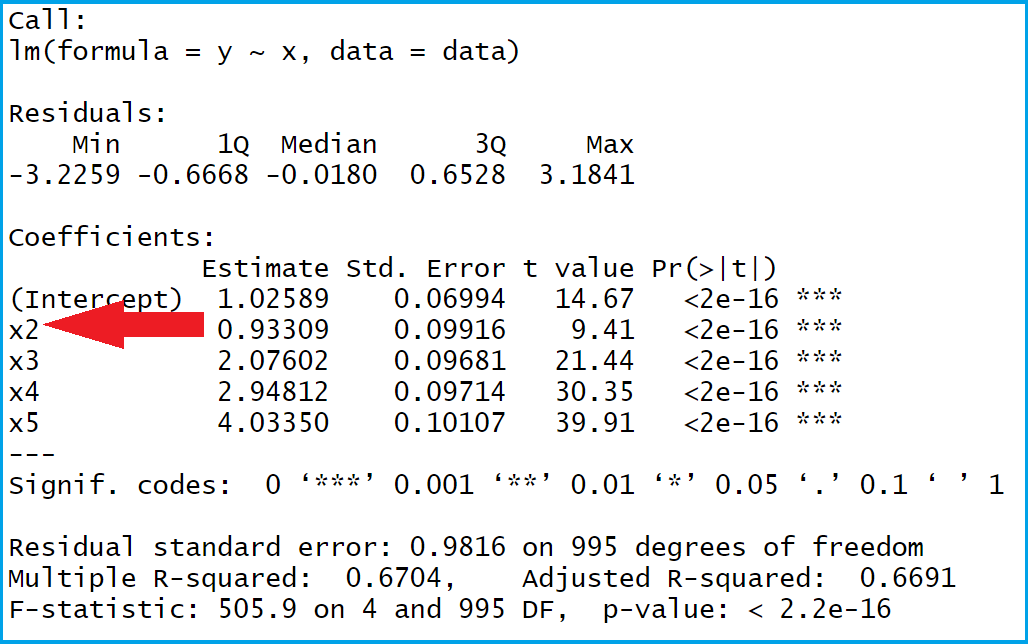 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
